Hello Brian! Tell me where you work and what you do.
I’m an assistant professor, soon to be an associate professor at the University of Georgia, in the Department of Plant Pathology. We study plant disease and plant immunity.
My lab, which I founded five and a half years ago, specifically works on different bacterial pathogens of plants. Right now we have 10 people in the lab, plus a few undergrad students who are short-term members.
We try to understand how bacterial pathogens subvert the immune system, as well as the mechanisms by which plants fight these pathogens.

Tell me about a challenge that your field faces.
We, humans, have an adaptive immune system — we produce antibodies, and we learn to defend against new types of pathogens. Plants can’t do that. Every germ and microbe that they’re ever going to be able to defend against is hard-encoded into their genes from the time they germinate to the time they die. So they do change, but they change on an evolutionary time-scale, or in the case of agriculture, a breeding time-scale.
My lab works on bacterial pathogens. A lot of plant pathogens are fungi, and for that, we have fungicide, which can be sprayed to remove the pathogens. But we don’t spray a field with antibiotics, it’s not appropriate or cost-effective to use for an entire field.
So, typically, to have effective management of bacterial plant diseases, you’re dependent upon genetic resistance. That creates challenges, because pathogens evolve quickly, and we can’t always breed plants fast enough. So if an entire field is made of genetic clones, you basically just need one pathogen that comes along and has the keys to wipe out the entire field.
What are you working on right now?
We have two projects going on right now.
For the first one, we’re working on the model of arabidopsis (it’s a model commonly used for genetics, but is also a good model host for disease). We’re currently focusing on one branch of the plant and its innate immune system that allows it to defend itself against the majority of microbes that it encounters, based on perception of conserved patterns. We’re trying to understand what it is about the immune response that actually shuts the pathogen down because that’s actually not clear. We understand a lot about the immune signaling side, how the plant cell responds to pathogens, and the different regulatory cascades involved, but we have a very poor understanding of what the output of this immune signaling is that actually controls pathogen growth.
We have sterilizing immunity - our body fully removes pathogens and maintains our internal spaces germ-free. But plants don’t. When plants have bacteria inside their leaves, which is a reasonable analog for our lungs (in terms of purpose and structure), the plants’ form of immunity keeps them from spreading disease but the bacteria isn’t going anywhere. They’re still there, still being maintained. It’s sort of a paradox — bacteria with nutrition, water and space, you’d expect them to grow. And yet, in the context of this environment, they don’t.
We’re trying to look at regulation of pathogens, genes, and different aspects of that from the pathogen side to understand what exactly the host is doing to those microbes. For this research, we’re supported by a grant from the national science foundation.
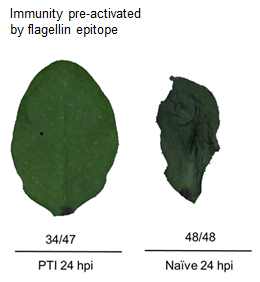

On the more applied side, we’ve been spending a lot of time and energy developing resources to understand pathogens of onion. Onion is a major commodity crop in our state. The vidalia sweet onion — a very low pungency, very tasty onion — is grown only in our part of the country. Onions are unusual in having a real menagerie of different bacterial pathogens that are of major concern.
We’re working on a relative of e-coli, called pantoea, and trying to understand how it causes disease, because when we looked at its genome it was missing a lot of the typical features you’d expect to see in a typical plant pathogen. We knew it was a pathogen but had no clues from its genetic sequence as to why that would be.
Over the years, we’ve identified some very important genes. This pathogen makes its own herbicide, and that’s a key function to their ability to cause disease — they literally kill the plant cells around them. So there’s no immune system anymore, they just live off the dead cells that they leave in their wake. Onions are an interesting crop for this strategy though, because the compounds that give onion its pungent taste are actual pretty potent antimicrobial compounds. So, the bacteria kills the cells and they release reactive sulfur compounds, and what we’ve identified is that it has a secondary pathway that allows it to specifically tolerate those chemicals, so it’s a kind of chemical warfare going on. Stacking these two forms together gives you a pretty unique form of disease, and that’s what we’ve been working on lately.
For this work we are supported by the US department of agriculture.
What is your role in the lab?
I’m the principal investigator. I’m the founder and head of the lab. I write all the grants, manage all the students, manage all the budget. Basically, I manage the research.
What are your biggest challenges as head of the lab?
Training new personnel, bringing in funds, and maintaining a steady output in the uncertain world of research. It’s hard to guarantee that a certain experiment will have results that lead to something new or interesting, it’s hard to coax information out of nature. And, arguably, our job is to turn money into information.
Why did you choose to start using Labguru?
When I got this job, I received a recommendation from another PI that I trusted. I saw that she was very organized with her research laboratory management and was handling a large group with a lot of complex projects, while using Labguru as a substitute for paper notebooks. It seemed like an interesting and reasonable solution, and having had to go through, in previous labs, the paper notebooks of previous lab members to try to interpret all of their experiments, I knew that a paper notebook can often be nearly useless outside of the hands of the person who wrote it. Labguru seemed like a solution to that problem.
Did it really prove to be a good solution to your problem?
Yes. With Labguru, we have useful digital notebooks that would maintain a good archive of data and interpretability. We also use it for other aspects of research laboratory management: maintaining our collections inventory, for example (it helps us make sure we keep track of our genetic inventory in a way that is accessible and everyone knows how to use it.)
Additionally, it helps with information preservation: with my first batch of students completing their doctorates and moving on, Labguru notebooks have helped protect and preserve the "institutional knowledge" in the lab.
How does Labguru answer your challenges as the head of the lab?
It lets me keep an eye on what people are doing, stay involved in their research. I can easily get key pieces of information from people’s data just by accessing their digital notes in Labguru. Having all of our group’s research at my fingertips has been a life saver for preparing manuscripts, reports, and presentations.
Does Labguru help with training students and new lab members?
Labguru provides a great platform for on-boarding. We use it to keep central documents to get people onboarded in the lab and direct them towards required safety training. We also use it to store our permit documents and SOPs so they are easily accessible by all lab members.
The lab notebook entries are built in a structured way that require you to enter information in the same fields every time, forcing people to think in a structured way. The organized, digital format is incredibly useful for new lab members to learn lab techniques or track down an experiment they need to repeat.
How do your lab members feel about Labguru?
My lab manager and lab members find labguru very valuable. They appreciate the structure and searchability. We are very invested in it and use it to store all our research information, and the inventory and sample management system serves us in our day-to-day research laboratory management actions.
There’s a very shallow learning curve with labguru that I’ve seen with people who join our lab, and once they’re committed to it they see the value of it.
Has Labguru helped you during the COVID-19 outbreak?
The same things that made Labguru useful in day-to-day work made it extra useful during the pandemic.
We could not use the lab simultaneously, so we would use Labguru to schedule shifts and let people know when they would or would not be using those shifts. This was an important part of our COVID-19 research resumption plan.
In addition, being able to access our data from any computer was invaluable when we were all at home.
I contacted a few of my colleagues requesting research information, and since they still use paper notebooks, I received answers such as “I can’t access that information, my lab notebook is in my lab, so I’ll get it to you in a couple of months.” That’s just not convenient. Unlike them, we could easily reach our information from any computer that’s connected to the internet.
Can you describe the ROI that you get from Labguru?
I'd estimate that on average we can find info on experiments at least twice as fast using Labguru as using paper notebooks, and it's 5 times as fast as digging through the notebook of a previous lab member. Each lab member probably saves 2 hours per week on average.
Would you recommend Labguru to other labs that are looking for a research laboratory management system?
If you’re starting a new lab or looking to make the switch over to an ELN and LIMS system, Labguru is definitely worth the investment.
To learn more about how Labguru can help you with research laboratory management, click here:


%20(4).png)